3/26—UK variant 61-64% more deadly
Increased mortality in community-tested cases of SARS-CoV-2 lineage B.1.1.7
Here we analyse a dataset linking 2,245,263 positive SARS-CoV-2 community tests and 17,452 COVID-19 deaths in England from 1 September 2020 to 14 February 2021. For 1,146,534 (51%) of these tests, the presence or absence of B.1.1.7 can be identified because of mutations in this lineage preventing PCR amplification of the spike gene target (S gene target failure, SGTF1). Based on 4,945 deaths with known SGTF status, we estimate that the hazard of death associated with SGTF is 55% (95% CI 39–72%) higher after adjustment for age, sex, ethnicity, deprivation, care home residence, local authority of residence and test date. Correcting for misclassification of SGTF and missingness in SGTF status, we estimate a 61% (42–82%) higher hazard of death associated with B.1.1.7. Our analysis suggests that B.1.1.7 is not only more transmissible than preexisting SARS-CoV-2 variants, but may also cause more severe illness.
Risk of mortality in patients infected with SARS-CoV-2 variant of concern 202012/1: matched cohort study
Participants were 54 906 matched pairs of participants who tested positive for SARS-CoV-2 in pillar 2 between 1 October 2020 and 29 January 2021, followed-up until 12 February 2021. The mortality hazard ratio associated with infection with VOC-202012/1 compared with infection with previously circulating variants was 1.64 (95% confidence interval 1.32 to 2.04) in patients who tested positive for covid-19 in the community. The probability that the risk of mortality is increased by infection with VOC-202012/01 is high. If this finding is generalisable to other populations, infection with VOC-202012/1 has the potential to cause substantial additional mortality compared with previously circulating variants.
An observational cohort study on the incidence of SARS-CoV-2 infection and B.1.1.7 variant infection in healthcare workers by antibody and vaccination status
[Preprint.] 13,109 HCWs participated; 8285 received the Pfizer-BioNTech vaccine (1407 two doses) and 2738 the Oxford-AstraZeneca vaccine (49 two doses). Compared to unvaccinated seronegative HCWs, natural immunity and two vaccination doses provided similar protection against symptomatic infection: no HCW vaccinated twice had symptomatic infection, and incidence was 98% lower in seropositive HCWs (adjusted incidence rate ratio 0.02 [95%CI <0.01-0.18]). Two vaccine doses or seropositivity reduced the incidence of any PCR-positive result with or without symptoms by 90% (0.10 [0.02-0.38]) and 85% (0.15 [0.08-0.26]) respectively. Single-dose vaccination reduced the incidence of symptomatic infection by 67% (0.33 [0.21-0.52]) and any PCR-positive result by 64% (0.36 [0.26-0.50]). There was no evidence of differences in immunity induced by natural infection and vaccination for infections with S-gene target failure and B.1.1.7. Natural infection resulting in detectable anti-spike antibodies and two vaccine doses both provide robust protection against SARS-CoV-2 infection, including against the B.1.1.7 variant.
Are vaccines safe in patients with Long COVID? A prospective observational study
[Preprint.] Forty-four vaccinated participants were assessed at a median of 32 days (IQR 20-41) post vaccination with 22 matched unvaccinated participants. Most were highly symptomatic of Long Covid at 8 months (82% in both groups had at least 1 persistent symptom), with fatigue (61%), breathlessness (50%) and insomnia (38%) predominating. There was no significant worsening in quality-of-life or mental wellbeing metrics pre versus post vaccination. Nearly two-thirds (n=27) reported transient (<72hr duration) systemic effects (including fever, myalgia and headache). Receipt of vaccination with either an mRNA or adenoviral vector vaccine was not associated with a worsening of Long Covid symptoms, quality of life, or mental wellbeing.
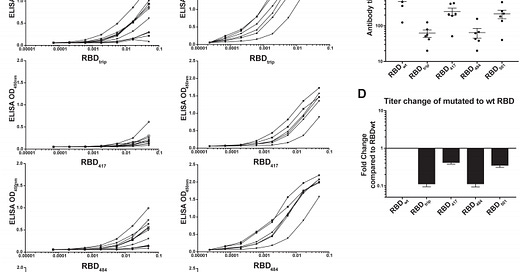
BNT162b2 mRNA COVID-19 vaccine induces antibodies of broader cross-reactivity than natural infection but recognition of mutant viruses is up to 10-fold reduced
[Preprint.] We aimed to assess how mutations in receptor binding domain (RBD) affected recognition of immune sera by antibodies induced by natural infection versus immunization with BNT162b2 [Pfizer vaccine]. We produced SARS-CoV-2 RBD mutants with single mutations in the receptor binding domain (RBD) region (E484K, K417N, N501Y) or with all 3 mutations combined, as occurring in the newly emerged variants B.1.351 (South Africa) and P.1 (Brazil). Our binding data demonstrate improved recognition of mutant viruses by BNT162b2-induced antibodies compared to those induced by natural infection. Recognition may, however, be 10-fold reduced for the variants B.1.351/P.1, suggesting that the development of a new vaccine is warranted.